The Evolutionary Genome Engineering Research Unit employs evolutionary genome engineering to accelerate laboratory evolution to combat antibiotic resistance and optimise biosynthetic processes, orders of magnitude faster than standard protocols. To this aim, we are developing in vivo mutagenesis methods with the ability to precisely generate vast genetic diversity in a wide range of microorganisms.
One of these developments, pORTMAGE (Nyerges, Á. et al. 2016. PNAS), enables the targeted construction of evolutionary trajectories in non-model species at an unprecedented speed. pORTMAGE is applicable to a continuously-expanding set of clinically and biotechnologically relevant bacterial species.
With our technologies:
- mutational processes that lead to novel protein-variants or better bio-synthetic pathways can be rapidly explored,
- evolutionary trajectories with multiple mutations can be mapped within a few days,
- the effects of clinically relevant antibiotic resistance-conferring mutations across bacterial pathogens and laboratory strains become comparable,
- the mode of action and the emergence of resistance against antibiotics can be better understood.
To this end, our research follows a DESIGN-BUILD-TEST cycle to mutagenize living cells and analyse evolutionary processes:
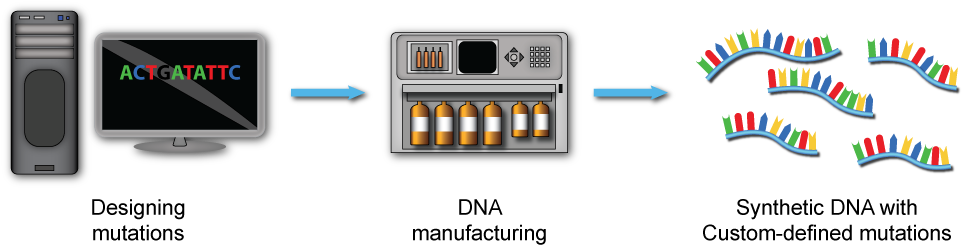
1. Design
We amalgamate functional genomics, system-level computational models, and adaptive laboratory evolution to gain greater insight into cellular mechanisms and evolutionary processes. We use Computer Aided DNA Design (CAD) and chemical DNA manufacturing to construct synthetic DNA molecules carrying predefined mutations.
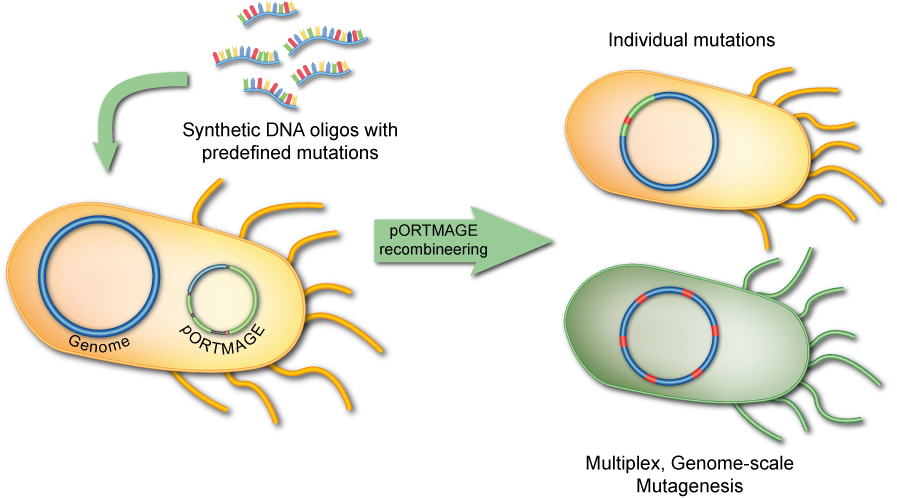
2. Build
pORTMAGE in conjunction with RNA-guided endonucleases (e.g. CRISPR) allow us to engineer cellular DNA in a diverse set of bacteria, including previously untapped, clinically- and biotechnologically-relevant species.
Key features of our technology:
- Plug-and-play genome engineering - mutagenizing, deleting or inserting features (mutations, genes, or entire biosynthetic pathways) into microbial genomes
- Efficient genome editing without off-target effects, in a diverse set of bacterial hosts with clinical- and biotechnological relevance
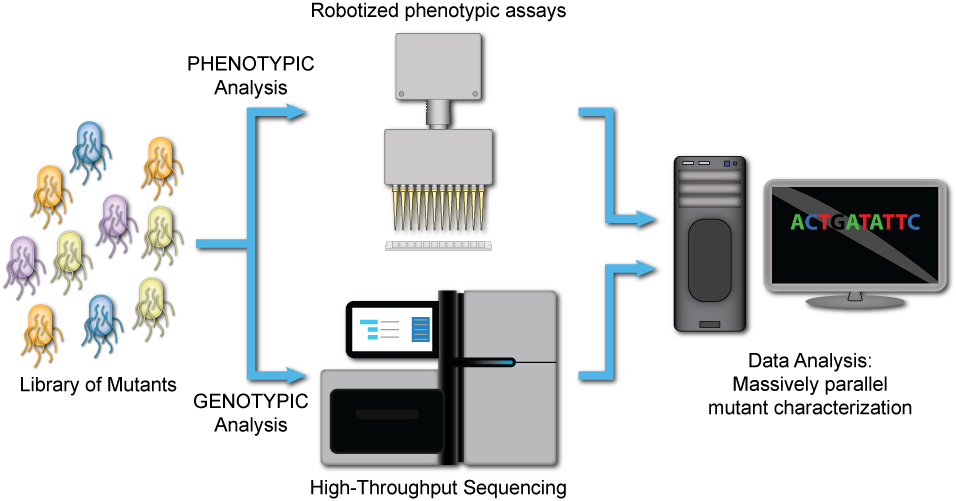
3. Test
Automated, high-throughput screening (HTS) and Next Generation Sequencing (NGS)-based genotype analysis allow us to link mutations to a phenotype.